Nifty way of detecting toxic strains of E. coli
Finding a fast and inexpensive way to detect specific strains of bacteria and viruses is critical to food safety, water quality, environmental protection and human health. However, current methods for detecting illness-causing strains of bacteria such as E. coli require either time-intensive biological cell cultures or DNA amplification approaches that rely on expensive laboratory equipment.
Now, Josh Hihath, an associate professor of electrical and computer engineering at the University of California, Davis, and colleagues at the University of Washington and TOBB University of Economics and Technology in Ankara, Turkey, have adapted a molecular electronic device called a single-molecule break junction to detect RNA from strains of E. coli known for causing illness. The findings were published in the journal Nature Nanotechnology.
“The reliable, efficient and inexpensive detection and identification of specific strains of microorganisms such as E. coli is a grand challenge in biology and the health sciences,” said Hihath. “Our technique could pave the way for rapid, straightforward detection of pathogens, antimicrobial-resistant bacterial strains and biomarkers for cancer.”
Hihath and his team focused on E. coli since it is a common pathogen that could easily be found in the food supply, but might not cause illness in a benign form. The worst strain of E. coli, called E. coli O157:H7, produces a toxic substance called Shiga toxin that causes bloody diarrhoea, kidney failure and even death.
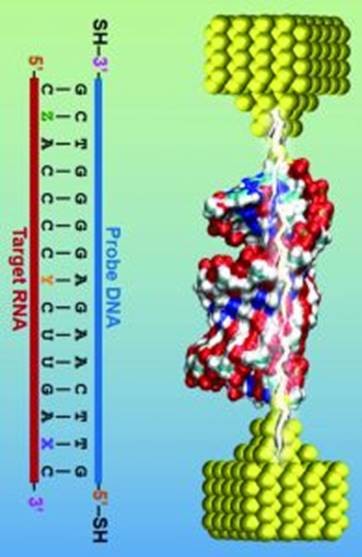
Single-molecule break junction devices consist of two metal electrodes with atomically sharp interfaces that are brought into contact in a liquid solution of interest, such as a solution containing RNA sequences from E. coli. As the electrodes are brought into contact and pulled apart, an electrical bias is applied and the current is measured. This process is repeated hundreds or thousands of times to determine the conductance of a single molecule.
“One of the questions we asked is, how small of a change in the sequence is needed to cause a meaningful change in the electrical conductance?” Hihath said. “The smallest thing we can change is a single-base, so we decided to see if a single-base change can be measured.”
By testing short sequences of RNA bound to DNA with chemical linkers, the team examined an E. coli sequence that would produce Shiga toxin. Their findings showed that changes in the electrical resistance of RNA due to a single-base change could be measured, which would allow them to see not only if a sequence was E.coli, but the specific strain of E.coli that produces Shiga toxin.
“A system that could selectively identify short sequences of DNA or RNA opens up new avenues for developing an electronic sensor platform for a wide range of applications,” he added. “Eventually, we want to get to the point where we can extract RNA samples from real organisms and measure their conductance on a sensing platform.”
The importance of bearing specification in bakery environments
Most commercial bakery environments are defined by temperature extremes, dust contamination and...
Weathering the storm: climate resilience in the food & beverage sector
As Australia's food and beverage manufacturing sector looks towards the near future, one...
Special K-ellanova upgrade for aging alarm system
Omniflex has delivered a direct retrofit solution for aging alarm systems at Kellanova’s...











